1D Linear operator with one parameter¶
This chapter introduces a basic example of the framework developed in Chapter 3. We take a one-dimensional system with a single parameter and extract an operator out of it.
It is trivial to verify linearity of the operator:
One of the solutions to this system might be:
:raw-latex:`\begin{align*} u(x) &= x^3 \\ f(x) &= \phi x^3 + 3x^2 \\ x &\in [0, 1] \end{align*}`
We define Gaussian priors on the input and output:
A noisy data model for the above system can be defined as:
For the sake of simplicity, we ignore the noise terms \(\epsilon_u\) and \(\epsilon_f\) while simulating the data. They’re nevertheless beneficial, when computing the negative log marginal likelihood (NLML) so that the resulting covariance matrix is mostly more well-behaved for reasons as they were outlined after the preface.
For the parameter estimation problem for the linear operator described above, we are given \(\{X_u, y_u\}\), \(\{X_f, y_f\}\) and we need to estimate \(\phi\).
Step 1: Simulate data¶
We use \(\phi = 2\).
In [2]:
def get_simulated_data(n1, n2, phi):
x_u = np.random.rand(n1)
y_u = np.power(x_u, 3)
x_f = np.random.rand(n2)
y_f = phi*np.power(x_f, 3) + 3*np.power(x_f,2)
return(x_u, y_u, x_f, y_f)
In [4]:
plt.show()
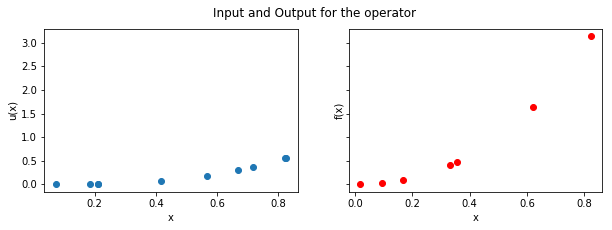
Step 2: Evaluate kernels¶
We use the RBF kernel defined as:
throughout the report. It is worth noting that this step uses information about \(\mathcal{L}_x^\phi\) but not about \(u(x)\) or \(f(x)\). The derivatives are computed using sympy.
In [5]:
x_i, x_j, theta, l, phi = sp.symbols('x_i x_j theta l phi')
kuu_sym = theta*sp.exp(-l*((x_i - x_j)**2))
kuu_fn = sp.lambdify((x_i, x_j, theta, l), kuu_sym, "numpy")
def kuu(x, theta, l):
k = np.zeros((x.size, x.size))
for i in range(x.size):
for j in range(x.size):
k[i,j] = kuu_fn(x[i], x[j], theta, l)
return k
In [6]:
kff_sym = phi**2*kuu_sym \
+ phi*sp.diff(kuu_sym, x_j) \
+ phi*sp.diff(kuu_sym, x_i) \
+ sp.diff(kuu_sym, x_j, x_i)
kff_fn = sp.lambdify((x_i, x_j, theta, l, phi), kff_sym, "numpy")
def kff(x, theta, l, phi):
k = np.zeros((x.size, x.size))
for i in range(x.size):
for j in range(x.size):
k[i,j] = kff_fn(x[i], x[j], theta, l, phi)
return k
In [7]:
kfu_sym = phi*kuu_sym + sp.diff(kuu_sym, x_i)
kfu_fn = sp.lambdify((x_i, x_j, theta, l, phi), kfu_sym, "numpy")
def kfu(x1, x2, theta, l, phi):
k = np.zeros((x2.size, x1.size))
for i in range(x2.size):
for j in range(x1.size):
k[i,j] = kfu_fn(x2[i], x1[j], theta, l, phi)
return k
In [8]:
def kuf(x1, x2, theta, l, phi):
return kfu(x1, x2, theta, l, phi).T
Step 3: Compute the negative log marginal likelihood(NLML)¶
The following covariance matrix is the result of our discussion at the end of Chapter 1.3.1, with an added noise parameter:
For simplicity, assume \(\sigma_u = \sigma_f\).
where \(y = \begin{bmatrix} y_u \\ y_f \end{bmatrix}\).
In [9]:
def nlml(params, x1, x2, y1, y2, s):
params = np.exp(params)
K = np.block([
[
kuu(x1, params[0], params[1]) + s*np.identity(x1.size),
kuf(x1, x2, params[0], params[1], params[2])
],
[
kfu(x1, x2, params[0], params[1], params[2]),
kff(x2, params[0], params[1], params[2]) + s*np.identity(x2.size)
]
])
y = np.concatenate((y1, y2))
val = 0.5*(np.log(abs(np.linalg.det(K))) \
+ np.mat(y) * np.linalg.inv(K) * np.mat(y).T)
return val.item(0)
Step 4: Optimize hyperparameters¶
In [10]:
nlml_wp = lambda params: nlml(params, x_u, x_f, y_u, y_f, 1e-6)
m = minimize(nlml_wp, np.random.rand(3), method="Nelder-Mead")
In [12]:
np.exp(m.x)
Out[12]:
array([11.90390211, 0.47469623, 2.00120508])
The estimated value comes very close to the actual value.
For the current model, we get the following optimal values of the hyperparameters:
| Parameter | Value |
|---|---|
| \(\theta\) | 11.90390211 |
| \(l\) | 0.47469623 |
| \(\phi\) | 2.00120508 |